Frictionless Linked Data
I was reminded of the importance of approachable, low-barrier-to-entry tools for data management by Monica Granados and Lily Zhao in their presentation of the Frictionless Data toolkit.1
They showcased use of a browser-based interface2 for a simple yet valuable task: associating title and description metadata with potentially cryptic column header names in a CSV file, and exporting that metadata together with the raw data to an open-source package format.
First, they directed the app to load an Algae.csv CSV file from an open data archive.3 The app
detected tabular data and offered to infer field names and data types:
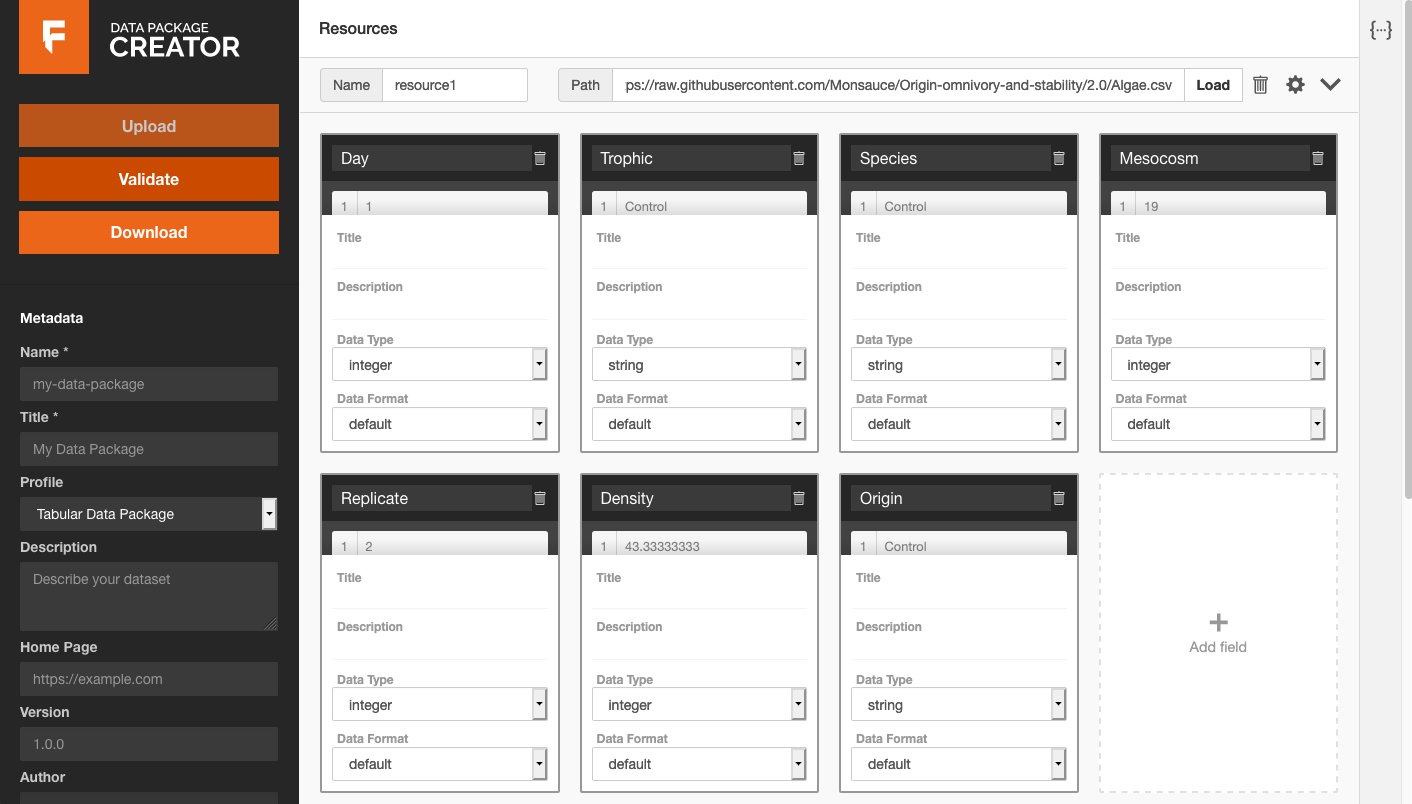
Next, they showed how a user could fill in field title and description metadata by hand, and select correct data types from a drop-down list:
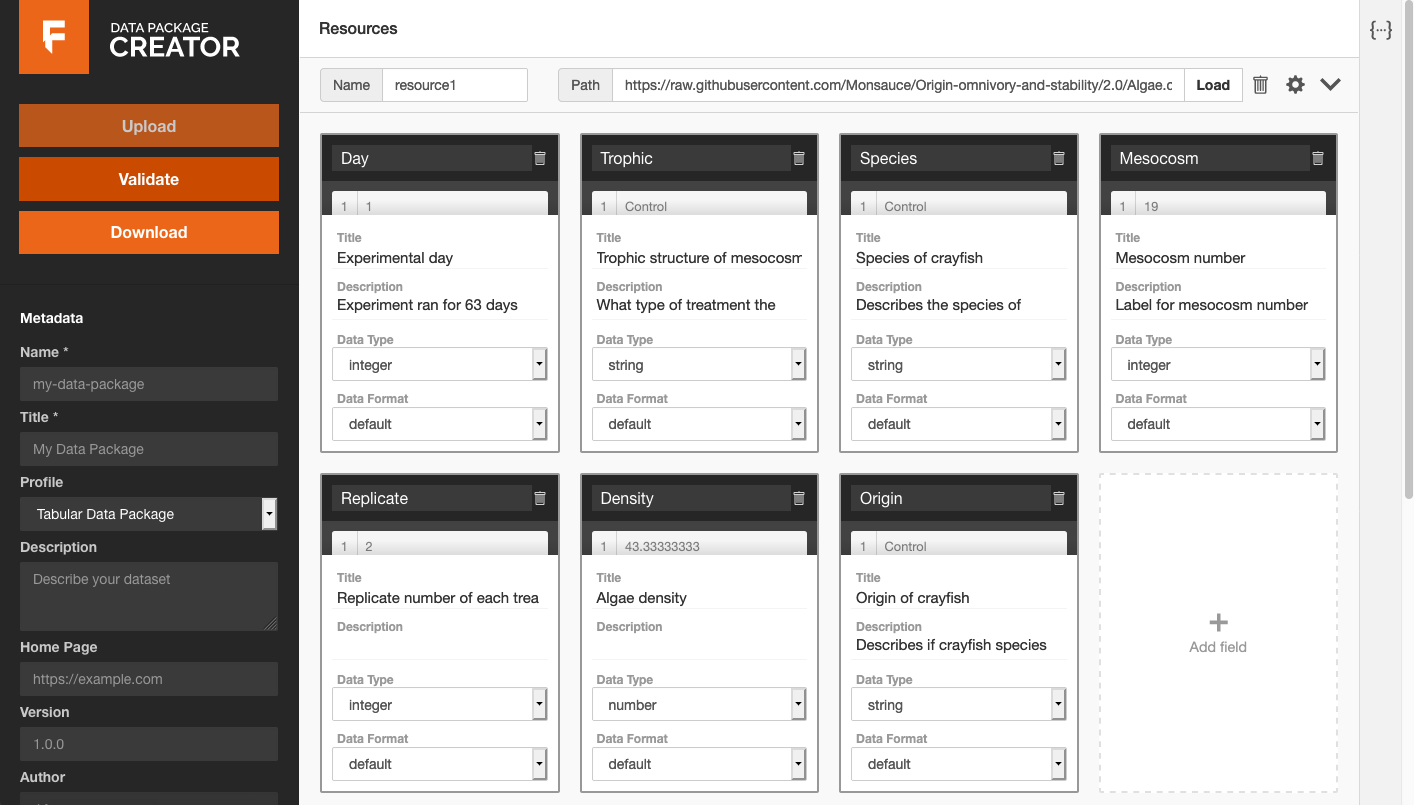
This kind of interface, where you are guided to fill in metadata and can produce shareable artifacts that are much more likely to be interpretable both by other humans and machines, is crucial for science that is more reproducible/trustable, and thus citable.
When I saw this demo, I immediately thought of the potential for linked open data to get this kind of
interface even closer to the Frictionless ideal. The Title and Description fields are analogous to
the rdfs:label and
rdfs:comment fields ubiquitous in linked open
data sets such as Wikidata, and there is
a rich vocabulary of data types as URIs under the namespace typically prefixed as
xsd:.
If user
input to Title and Description were to be fuzzy-matched to corresponding linked-data field values,
the app could auto-suggest additional metadata for the tabular data column. Unambiguous
URIs from Wikidata or other sources could be associated as metadata, making it even more likely
that the recipient of a data package could understand the semantics of the data – by following links.
References
Granados and Zhao, “How Frictionless Data can help you grease your data”. csv,conf,v5 conference, 2020. ↩︎
Data Package Creator, https://create.frictionlessdata.io/. ↩︎